pET-27b
Search name
pET-27b,Plasmid pET-27b,pET-27b vector
pET-27b Informaiton
Promoter: T7/lac
Replicon: ColE1 ori, F1 ori
Terminator: T7 Terminator
Plasmid classification: Escherichia coli plasmid; large intestine expression plasmid; pET series plasmid.
Plasmid size: 5414bp
Plasmid tagging: N-pelB, C-6 x His, C-HSV
Prokaryotic resistance: kanamycin Kan (50 g/ml)
Cloning strains: E. coli DH5 and E.
Culture conditions: 37 C, aerobic, LB
Expression host: E. coli BL21 (DE3).
Culture conditions: 37 C, aerobic, LB
Induction: IPTG or lactose and its analogues.
5'sequencing primers: T7: (TAATACGACTCACTATAGGG)
3'sequencing primers: T7-ter: (TGCTAGTTATTGCTCAGCGG)
Remarks: secreted expression
pET-27b Description
pET-27b is a prokaryotic expression vector. The C-terminal contains a 6*His tag and a HSV tag. The N-terminal contains a pelB precursor peptide. The plasmid contains several commonly used restriction sites to facilitate cloning of different genes. The expression was induced by T7 RNA polymerase from host cells. The target gene was cloned into plasmid vector and controlled by strong phage transcription and translation signals.
pET system is the most powerful system ever used to express recombinant protein in E. coli. It is also the most widely used system in prokaryotic expression. The plasmids can easily reduce protein expression by decreasing the concentration of inducers. Under non inducible conditions, the target gene can be completely silenced without transcribing.
pET-27b(+) vector carries an N-terminal pelB signal sequence for potential periplasmic localization, plus optional C-terminal HSV•Tag® and His•Tag® sequences. This vector differs from pET-25b(+) only by its selectable marker (kanamycin vs. ampicillin resistance).Unique sites are shown on the circle map. Note that the sequence is numbered by the pBR322 convention, so the T7 expression region is reversed on the circular map. The cloning/expression region of the coding strand transcribed by T7 RNA polymerase is shown below. The f1 origin is oriented so that infection with helper phage will produce virions containing single-stranded DNA that corresponds to the coding strand. Therefore, single-stranded sequencing should be performed using the T7 terminator primer .
pET-27b Multiple cloning site
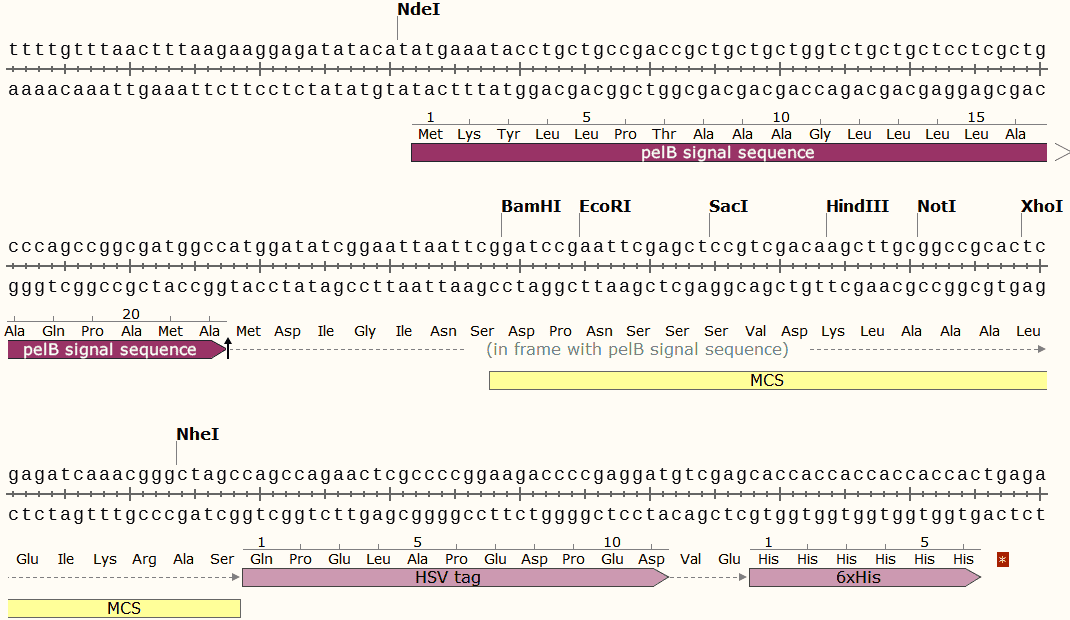
pET-27b Sequence
LOCUS Exported 5414 bp ds-DNA circular SYN 15-JUN-2017
DEFINITION synthetic circular DNA
KEYWORDS pET-27b
SOURCE synthetic DNA construct
ORGANISM synthetic DNA construct
REFERENCE 1 (bases 1 to 5414)
TITLE Direct Submission
JOURNAL Exported Thursday, June 15, 2017
FEATURES Location/Qualifiers
source 1..5414
/organism="synthetic DNA construct"
/mol_type="other DNA"
rep_origin 12..467
/direction=RIGHT
/note="f1 ori"
/note="f1 bacteriophage origin of replication; arrow
indicates direction of (+) strand synthesis"
CDS complement(560..1375)
/codon_start=1
/gene="aph(3')-Ia"
/product="aminoglycoside phosphotransferase"
/note="KanR"
/note="confers resistance to kanamycin"
/translation="MSHIQRETSCSRPRLNSNMDADLYGYKWARDNVGQSGATIYRLYG
KPDAPELFLKHGKGSVANDVTDEMVRLNWLTEFMPLPTIKHFIRTPDDAWLLTTAIPGK
TAFQVLEEYPDSGENIVDALAVFLRRLHSIPVCNCPFNSDRVFRLAQAQSRMNNGLVDA
SDFDDERNGWPVEQVWKEMHKLLPFSPDSVVTHGDFSLDNLIFDEGKLIGCIDVGRVGI
ADRYQDLAILWNCLGEFSPSLQKRLFQKYGIDNPDMNKLQFHLMLDEFF"
rep_origin 1497..2085
/direction=RIGHT
/note="ori"
/note="high-copy-number colE1/pMB1/pBR322/pUC origin of
replication"
CDS complement(2515..2706)
/codon_start=1
/gene="rop"
/product="Rop protein"
/note="rop"
/translation="MTKQEKTALNMARFIRSQTLTLLEKLNELDADEQADICESLHDHA
DELYRSCLARFGDDGENL"
CDS complement(3515..4597)
/codon_start=1
/gene="lacI"
/product="lac repressor"
/note="lacI"
/note="The lac repressor binds to the lac operator to
inhibit transcription in E. coli. This inhibition can be
relieved by adding lactose or
isopropyl-beta-D-thiogalactopyranoside (IPTG)."
/translation="MKPVTLYDVAEYAGVSYQTVSRVVNQASHVSAKTREKVEAAMAEL
NYIPNRVAQQLAGKQSLLIGVATSSLALHAPSQIVAAIKSRADQLGASVVVSMVERSGV
EACKAAVHNLLAQRVSGLIINYPLDDQDAIAVEAACTNVPALFLDVSDQTPINSIIFSH
EDGTRLGVEHLVALGHQQIALLAGPLSSVSARLRLAGWHKYLTRNQIQPIAEREGDWSA
MSGFQQTMQMLNEGIVPTAMLVANDQMALGAMRAITESGLRVGADISVVGYDDTEDSSC
YIPPLTTIKQDFRLLGQTSVDRLLQLSQGQAVKGNQLLPVSLVKRKTTLAPNTQTASPR
ALADSLMQLARQVSRLESGQ"
promoter complement(4598..4675)
/gene="lacI"
/note="lacI promoter"
promoter 4984..5002
/note="T7 promoter"
/note="promoter for bacteriophage T7 RNA polymerase"
protein_bind 5003..5027
/bound_moiety="lac repressor encoded by lacI"
/note="lac operator"
/note="The lac repressor binds to the lac operator to
inhibit transcription in E. coli. This inhibition can be
relieved by adding lactose or
isopropyl-beta-D-thiogalactopyranoside (IPTG)."
CDS 5072..5137
/codon_start=1
/gene="pelB (fragment)"
/product="leader peptide for secretion"
/note="pelB signal sequence"
/translation="MKYLLPTAAAGLLLLAAQPAMA"
CDS 5219..5251
/codon_start=1
/product="HSV (herpes simplex virus) epitope tag"
/note="HSV tag"
/translation="QPELAPEDPED"
CDS 5258..5275
/codon_start=1
/product="6xHis affinity tag"
/note="6xHis"
/translation="HHHHHH"
terminator 5342..5389
/note="T7 terminator"
/note="transcription terminator for bacteriophage T7 RNA
polymerase"
ORIGIN
1 tggcgaatgg gacgcgccct gtagcggcgc attaagcgcg gcgggtgtgg tggttacgcg
61 cagcgtgacc gctacacttg ccagcgccct agcgcccgct cctttcgctt tcttcccttc
121 ctttctcgcc acgttcgccg gctttccccg tcaagctcta aatcgggggc tccctttagg
181 gttccgattt agtgctttac ggcacctcga ccccaaaaaa cttgattagg gtgatggttc
241 acgtagtggg ccatcgccct gatagacggt ttttcgccct ttgacgttgg agtccacgtt
301 ctttaatagt ggactcttgt tccaaactgg aacaacactc aaccctatct cggtctattc